In a previous article we introduced a "A Simple DSLR Camera Sensor Noise Model" and then we used this model to analyze a camera sensor raw noise in "Use of Linear Regressions to Model Camera Sensor Noise". Those articles deal with the theoretical and practical basis to analyze noise from raw image samples.
The input to that analysis are samples of photographs taken at different exposition levels, from which we compute the variance and mean of the photosite readings. Until now, we have prepared the samples and computed their variance and mean using Iris Software. But that workflow was very labor intensive, error prone, and inhibit us to further analysis like the study of the covariance between channels in the same sample because Iris software does not bring that information. Furthermore, Iris is a windows only software, which prevents users with other OSs to collect noise data using this tool.
To solve those issues, we have prepared —and still is a work in progress— a R package called imgnoiser (from IMaGe NOISE R package). In this article, we introduce you to its usage, which automatizes the acquisition, analysis noise statistics, letting us to focus in the study, instead of devoting a lot of time to collect the data or to prepare code to study that data.
Without the use of this package I could spent a whole morning just collecting the noise statistics. Now, with the help of imgnoiser it takes me only few minutes and brings additional and more accurate data than Iris.
You can install the package from github using:
devtools::install_github('oscardelama/imgnoiser')
In the imgnoiser repository wiki, in github, you will also find the package vignettes including the list of the hvdvm and vvm classes components, along with their functions and its parameters.
If you install this package, you will have in your own R session, detailed documentation of the package functions, their parameters, their default and possible values, the package options and so on. All of that is, or eventually will be, available in your R help system.
This article explains general concepts and the package workflow usage with "real life" examples. In that sense, we will mention the main functions, their main parameters and so on, but the minute technical detail of all the functions and their arguments is not the focus of this post; please read about that in the R help system or the package vignettes.
If using this package you find any issue please file it here.
In the R code excerpts, the function output or messages are prefixed with #> while the real comments are prefixed just with #.
Reproducible Research
This package is built having in mind the reproducible research concept. To make a research reproducible, all the code and data to reproduce the research figures must be delivered, including a data dictionary and an explanation of the complete software development environment, its configuration and the whole set of instructions which generated each intermediate or final tables and images.
The focus of imgnoiser is to factorize most of that software environment, configuration, data definitions and conventions in one single point, to easily share and reuse noise analysis papers without the need to include, define and explain repeatedly the software environment and instructions to reproduce most of the results.
Collecting the Image Samples
Before the existence of the imgnoiser package, after taking the photographs for the noise analysis, we cropped from them the region of our interest (ROI) using Iris Software. Now we can use dcraw. Dcraw is a very well known open source and free software product, with its own wikipedia entry, capable to develop raw image files from most of the DSLR camera brands an models. Many open raw photo decoders use dcraw under the hood. For example (quoted from the dcraw web page), ACDSee, Adobe Photoshop, Picasa from Google and Rawtherapee among many others.
Dcraw is developed and maintained by David Coffin using Ansi C code, this make it very portable to any OS. I am not very familiar with Linux, but I understand it can be compiled in just one command line, please check the dcraw web site, which explains that.
You can find in the web the way to get the binaries in other OSs. For example, this page explains how to compile dcraw for OS X Mavericks, and in this other one you can find a link to download the windows executable for 32 and 64-bits systems, this is provided by Axel Rietschin, which is a trustworthy source, and is the version I will use for this post in a Windows 7, 64-bits system.
The way to take the photographs to get the image samples is explained here and also in a vignette in the imgnoiser package that you can read here. We will assume you have just taken the photographs and you want to know how to handle them in order to process the samples with imgnoiser.
The Package Options
The imgnoiser package contains a set of options with factory settings you can change to customize the package at your taste.
For example the plotting color palette 'color.pallette' and the default RGGB channel labels 'RGGB.channel.labels' are among those options. Very often, they are used by the package functions as default values. This way you can fit those options and use less arguments while calling those functions.
Calling the imgnoiser.option() function without arguments, you will get a list of all the package parameters. If you call it with the name of an option you will get its value, or NULL if it doesn't exists. If you call it with an option name and a value you will set that option with that value.
# Get the complete list of package options
imgnoiser.option()
#> $fit.model.family
#> [1] "lm"
#>
#> $fit.model.data
#> [1] "std.var"
#>
#> $fit.model.name
#> [1] "standard"
#>
#> <Option list truncated here>
# Get the current confidence level option
imgnoiser.option('conf.level')
#> [1] 0.95
# Set the confidence level to 98%
imgnoiser.option('conf.level', 0.98)
# Check the new value
imgnoiser.option('conf.level')
#> [1] 0.98
You can save the package options and load them in a different R session. Use the imgnoiser.options.save() function to save the options and imgnoiser.options.load() to load them. Check the R help system for the details.
Getting the Raw Image Samples
Dcraw is mostly used to process the raw file and get the final .jpg or .tif image. However, we want it to extract the raw image without any change of this raw data in the process. To get that we just have to use the -D -4 command options. For example, I will open a command line window and set the folder containing the .NEF (Nikon raw format) photographs I want to process, as the current working directory: you can shift+right+click on the folder in the Windows file Explorer and choose Open command window here from the popup menu that appears. Then I use:
for %f in (*.NEF) do dcraw -D -4 %f
Previously, I have set the path to the dcraw executable in my system PATH environment variable. If you don't wan't or don't know how to do that, just prefix dcraw with the path to the 'dcraw.exe' file, as in ... do C:\path\to\dcraw -D ....
We can't just use dcraw -D -4 *.NEF because dcraw does not understand Windows wildcard characters like *. If you want to put that command line in a batch file, you must to use %%f instead %f.
Now my folder with the .NEF files also contains .pgm ones, containing the raw images. The .pgm format is an open format and stands for Portable Gray Map. In opposition to .NEF, this is a format that imgnoser can read.
For each processed .NEF file now there is a .pgm one with the same base name as in:
05/11/2013 12:39 p.m. 16,331,695 _ODL5762.NEF
08/01/2015 05:58 p.m. 32,458,899 _ODL5762.pgm
Now we should find the region of interest (ROI) in the images to later crop them from each photograph. To do that we should pick a "well exposed" image, this means one with the image data in the third quartile along the horizontal axis in the image histogram. We are looking for the area of the image most evenly lit, otherwise we will measure variance caused by shadows in the image instead of noise variance.
Remember the raw image is composed by only one layer (there are not red , green and blue layers as in RGB images), like in a gray image, where the pixels follow a 2x2 color filter (channels) array pattern. First we will separate those four channels using the imgnoiser::split_channels function:
path <- 'path/to/samples'
cfa <- imgnoiser::split_channels('_ODL5719.pgm', path)
str(cfa)
# List of 4
# $ ch1: int [1:1640, 1:2474] 962 986 948 934 949 1008 944 959 984 961 ...
# $ ch2: int [1:1640, 1:2474] 1940 1998 1983 1984 1972 1980 1975 2033 1993 1992 ...
# $ ch3: int [1:1640, 1:2474] 1963 2015 1997 1990 1994 2035 1982 1955 2034 2027 ...
# $ ch4: int [1:1640, 1:2474] 1469 1437 1472 1527 1479 1528 1477 1502 1450 1524 ...
The above list of channels, are labeled according to 2x2 channel pattern in the image starting from the image top left corner:
+-----+-----+---
| ch1 | ch2 |
+-----+-----+---
| ch3 | ch4 |
+-----------+---
|
Most of the cameras have the RGGB pattern array. This way (ch1, ch2, ch3, ch4) == (red, greenR, greenB, blue), where greenR stands for green in the red row and —naturally— greenB stands for green in the blue row. If you are not sure if that is the case for your camera, remember that for a white target the channels with highest and similar values correspond to the green channels, then the blue and finally the red one. If you look the values in the result from the R code above, it is clear they have the RGGB pattern.
Now we will save one of the green channels using the writeTIFF function in the tiff package. That function expects values in the [0,1] range. Accordingly, we will divide each pixel value in the greenR channel by 16,383 ADU which is the maximum possible in the 14-bit .NEF format.
library(tiff)
matx <- cfa$ch2 / 16383
tiff::writeTIFF(matx, 'green_r.tif', bits.per.sample = 16L, compression='LZW')
# [1] 1
Now we have a .tif image which we open using ImageJ Software. We will remove pixel values variance caused by noise to get the overall illumination in the image. Using the FFT bandpass filter we will keep the image features with a size above 6 pixels (menu command "Process » FFT » Bandpass Filter..."), then we will see the resulting image using the menu command "Plugins » 3D » Interactive 3D Surface Plot", with this tool we can interactively rotate and scale the rendered surface, where its elevation represents the light intensity.
It is clear the reddish area is the more evenly lit region. The holes in the surface are dirty spots in the camera sensor.
Using this 3D view tool, we will look for a region, of at least 32x32 pixels, evenly lit and without including a dirt spot. Proceeding that way, I select a rectangle as the best region, and find out this corresponds to an area of 144x114 px with the top left corner at (496,726). From this area, finally I choose a 40x40 px square (1600 pixels) at (496, 726).
Notice we are working with pixels from one channel only, whose dimensions are half the ones of the whole image. To map this ROI in the channel to the whole image we have to multiply the coordinates and the region dimensions by 2. This way we get a 80x80 pixels square with its top left corner at (992, 1452).
If you really want to find the most evenly lit area in your shots, after the bandpass filter described above, you can convert the resulting image to floating point pixel values with the menu command "Image » Type » 32-bit", duplicate that image and apply "Process » Filter » Variance" with 40 pixel radius, then take the square root to that image to get the standard deviation ("Process » Math » Square Root") and divide it by the other bandpass copy, to get the relative standard deviation. Set the brightness and contrast to a proper image presentation ("Image » Adjust » Brightness/Contrast"). Now look for the darkest and flattest area with at least 32x32 pixels, look how flat it is with the 3D viewer tool.
In other article I mentioned finding an area with the smaller variance, but the issue with that technique is that the darkest areas get a lower variances without being necessarily more uniformly lit, computing the relative standard deviation as indicated above, counterbalance that fact.
The coordinate system origin in ImageJ is the (0,0), corresponding to the pixel at the image top left corner. This means that if we choose both coordinates with even values, the samples will also have the RGGB pattern. This is very important to do not later get confused because the samples have an unexpected channel pattern. It is also important to choose an area with even dimensions, that way we will have in each sample the same quantity of pixels from each channel.
Now we have to clip this region from each of the photographs. To that effect we will use an Image Magick tool. This is also free software, available for Linux, Windows and Mac OS. To clip our ROI, from the raw images we got from dcraw, we have to execute the mogrify command this way:
mogrify -extract <width>x<height>+<horiz pos>+<vert pos> <image file name>
Where <width> and <height> are the clipping dimensions, and
mogrify -extract 40x40+992+1452 _ODL*.pgm
The files resulting from this command, replace the the original ones. Instead of the set of big (32MB) .pgm files, now I have a set of small 4KB files:
08/01/2015 09:40 p.m. 3,215 _ODL5752.pgm
08/01/2015 09:40 p.m. 3,215 _ODL5753.pgm
If you want to keep your original, big .pgm files, perhaps to use another image area as an additional set to your samples, you can use the path option in the mogrify command. For example, if you add the option -path crops the resulting crops will be saved in the crops sub-folder, leaving your original .pgm intact.
Now we have the sample images ready to be processed by imgnoiser, you can download them from here (134 KB). Notice that except of locating the ROI, the required activities are just a couple of command lines that take just a few seconds of processing.
Brief Introduction to the Usage of R6 Classes
Some of the imgnoiser functions are organized in R6 classes, which means its usage is slightly different to other R packages. However, I am sure you will find this more convenient. In this section we will briefly learn the general usage of this classes. What will see is applicable to any R6 class not just those in imgnoiser.
The first step first to be able to access to the services in a class is the creation of an instance of that class. This instance is also called class instance object or just object for short.
Typically the object creation is done through the use of the new function. For example, the imgnoiser package contains the vvm class. So, we can create an instance object of the vvm class using:
my.object <- vvm$new(<your creation parameters>)
Now, the variable my.object contains an object of the vvm class, which we will use to access to the class services. During the usage of this object, it will carry, inside itself, the information resulting from the functions we will call using the object. For example, lets suppose we have already processed some image samples and obtained some noise statistics about them, and now we want to fit a linear model over those statistics. At this point in time, my.object will already contain the image samples statistics and we can fit the model just using:
my.object$fit.model(<your model fitting parameters>)
The point here is, we don't have to keep R variables holding the image samples statistics. That information is kept inside my.object, so we can just call the fit.model() function without the need to provide that information as arguments in that call. Also, the result of the model fitting is now also inside the object. Again, we don't need to keep those fitted models in other R variables.
As a final example, lets suppose we want to plot the image samples statistics and the predictions from the fitted model. As the fitted model, and the image statistics, are kept inside the object, we can just call the plotting function with arguments focused only on the plot. i.e. We don't have to provide the statistics and the fitted model as arguments to the plotting function.
my.object$plot(<your plotting parameters>)
As you may see, this is very convenient and more easier to use than continuously providing the different input data required by the class functions and saving their results.
Notice that even when fit.model and plot are functions of the vvm class (technically they are methods) we call them using the object we created at the very beginning. That makes sense considering each object holds the data produced in its previous usage.
A class can also contain variables. For example, the vvm class includes the var.df variable, containing a data frame with the noise variance information. As usual, we access to that class component through an object, my.object in our examples. As with any R variable, we can read the value in the object variable, for example, to pass it as argument to other function or to set other variable, just by itself or as part of an expression:
my.var <- my.object$var.df
other_var <- subset(my.object$var.df, <my subset conditions>)
some_function(..., my.object$var.df)
In the imgnoiser package most of those variables are read-only. However, technically its is possible to write or set values on them, eventually we might end up having some of them.
my.object$some.variable <- some_value
You can print the instance object to get a list of its components to remember what you can do with it.
my.object
#> <vvm>
#> function:
#> digest()
#> digest.as.rgb()
#> digest.from.rgb()
#> exists.model()
#> fit.model()
#> get.model()
#> get.model.predictions()
#> initialize()
#> plot()
#> print.model.summary()
#> remove.model()
#> variable:
#> channel.labels
#> cov.df
#> green.channels
#> merged.var.cov.df
#> model.list
#> RGGB.indices
#> var.df
#> wide.var.df
To get detailed documentation about this components, you can use the R help system, for example:
help('vvm$fit.model')
Digesting the image Samples
There are two way to process the image samples, each one is represented by one class in the package, the vvm and the hvdvm class. In both classes the function that process the raw image samples is named digest(). Except for the way how they process the samples, both are very similar. However, each way of processing brings different output, in their values as in their meaning.
Digesting Samples with the vvm Class
The vvm class computes the regular mean, variance and covariance in the image samples, expecting a quadratic relationship between the pixel values variance and their mean. The theoretical basis of this approach is explained here and the practical application of that model is explained here.
The vvm class can digest the raw image samples in more than one way. In the simplest way it process the data as-is in the raw image samples. However, this class can convert the raw images to a RGB space an the process the resulting RGB samples, this help us to track the image noise from its origin in the raw image to the final sRGB or Adobe RGB profile.
Digesting the Samples as Raw Data
We will process the samples with the vvm class. Lets create a vvm object with name vvm.obj. During the instance creation we can specify the channel names, which of those are the green channels and so on. The easiest way to do all that, when the samples have the RGGB pattern —as in our case— is with the parameter has.RGGB.pattern set to TRUE.
vvm.obj <- vvm$new(has.RGGB.pattern = TRUE)
In the call to the digest() function we will specify the samples we want to get processed by giving the alphabetical range of their base name, their path and their file name extension. In the following example we want to process all the images between (and including) the files from 'ODL5719.pgm' to 'ODL5767.pgm' in the './samples/' folder.
vvm.obj$digest(file.name.from = '_ODL5695',
file.name.to = '_ODL5767',
file.name.ext = '.pgm',
file.path = './samples/')
A progress bar will appear while the function is crunching the samples, it takes approximately 5 seconds the processing of 200 samples.
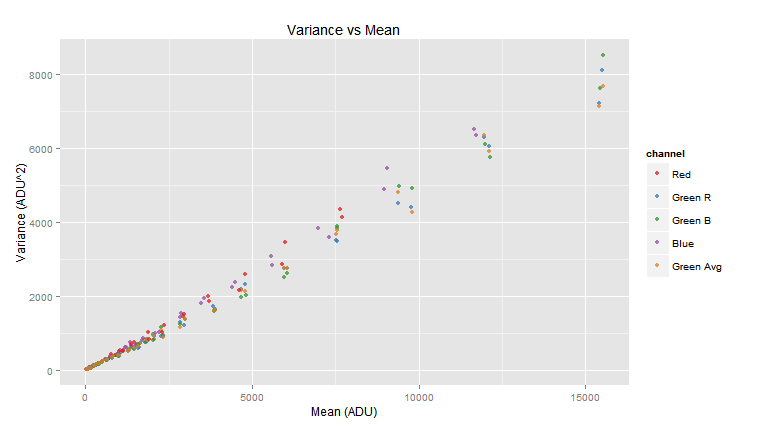
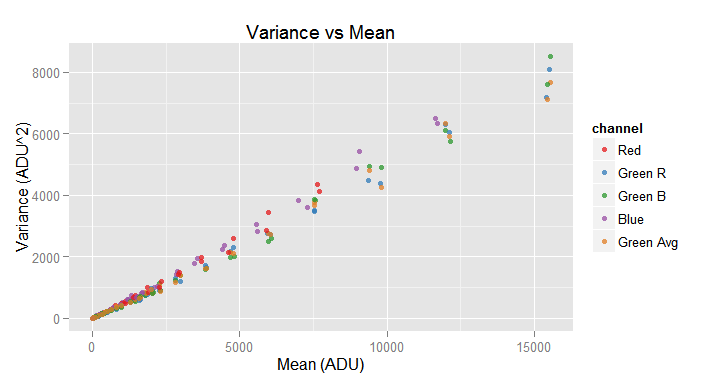
Notice that just by calling three functions we have done all the noise data collection! Now we can sit back and take a look to the plot of the variance against the mean for each channel in the image samples.
vvm.obj$plot()
You may notice, in the above graph legend, a channel labeled Green Avg this "channel" shows the values averaging both green channels. In a later section we will see in detail the plot() function.
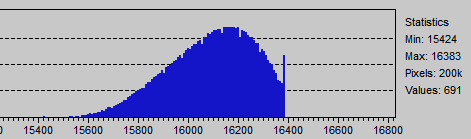
The following image shows the kind of image sample we don't want to include in the set of digested samples.
A partially or totally clipped sample have smaller variance than it should have if such clipping would not occurred, also their mean value is distorted.
Of course it is not possible to check always each sample histogram to discard these cases. The digest() function parameters min.raw and max.raw helps to detect this situation. If more than the 2% of the pixel values in a channel sample is outside the given [min, max] range, the sample is discarded. This parameters also help us to process only the noise in a particular segment of its whole range, for example close to the origin.
These arguments must be numeric vectors, each with one or four values. If only one value is given, the same one is used for all the channels, otherwise each channel have its own minimum or maximum value.
To get the data computed by the digest() function, part of which is shown in the plot above, we can check the var.df and the cov.df object variables. They are data frames containing —correspondingly— the variance and covariance data collected from the images samples.
head(vvm.obj$var.df, 8)
| pict | channel | mean | var |
|---|---|---|---|
| _ODL5695.pgm | Red | 696.1 | 314.5 |
| _ODL5695.pgm | Green R | 1410.8 | 618.1 |
| _ODL5695.pgm | Green B | 1414.6 | 597.3 |
| _ODL5695.pgm | Blue | 1033.9 | 518.7 |
| _ODL5695.pgm | Green Avg | 1412.7 | 593.5 |
| _ODL5696.pgm | Red | 716.3 | 322.2 |
| _ODL5696.pgm | Green R | 1453.1 | 624.6 |
| _ODL5696.pgm | Green B | 1456.2 | 566.8 |
head(vvm.obj$cov.df, 8)
| pict | chan.a | chan.b | cov.a.b |
|---|---|---|---|
| _ODL5695.pgm | Red | Green R | 0.3391 |
| _ODL5695.pgm | Red | Green B | 36.1396 |
| _ODL5695.pgm | Red | Blue | 16.0167 |
| _ODL5695.pgm | Red | Green Avg | 18.2393 |
| _ODL5695.pgm | Green R | Green B | -14.2368 |
| _ODL5695.pgm | Green R | Blue | 4.7311 |
| _ODL5695.pgm | Green R | Green Avg | 301.9246 |
| _ODL5695.pgm | Green B | Blue | -19.0844 |
Notice those data frames are in a rather narrow form. A sign of that is they use more than one row for data from the same image sample. In the case of the var.df data, you can get it in a wide form using the wide.var.df.
head(vvm.obj$wide.var.df, 8)
| pict | red.mean | red.var | green.r.mean | green.r.var | green.b.mean | green.b.var | blue.mean | blue.var | green.avg.mean | green.avg.var |
|---|---|---|---|---|---|---|---|---|---|---|
| _ODL5695.pgm | 696.1 | 314.5 | 1411 | 618.1 | 1415 | 597.3 | 1034 | 518.7 | 1413 | 593.5 |
| _ODL5696.pgm | 716.3 | 322.2 | 1453 | 624.6 | 1456 | 566.8 | 1063 | 479.0 | 1455 | 599.7 |
| _ODL5697.pgm | 882.7 | 391.2 | 1800 | 743.2 | 1802 | 740.7 | 1329 | 740.5 | 1801 | 834.2 |
| _ODL5698.pgm | 906.2 | 394.9 | 1852 | 767.3 | 1850 | 818.2 | 1364 | 652.4 | 1851 | 809.2 |
| _ODL5699.pgm | 1112.7 | 544.9 | 2276 | 901.3 | 2278 | 1142.6 | 1682 | 796.6 | 2277 | 1072.0 |
| _ODL5700.pgm | 1136.3 | 496.4 | 2317 | 923.4 | 2321 | 924.1 | 1713 | 847.2 | 2319 | 881.3 |
| _ODL5701.pgm | 1381.6 | 684.7 | 2830 | 1285.7 | 2833 | 1246.3 | 2097 | 996.4 | 2832 | 1164.3 |
| _ODL5702.pgm | 1452.9 | 748.7 | 2973 | 1206.1 | 2978 | 1381.8 | 2203 | 1027.7 | 2976 | 1381.7 |
This table contains exactly the same data as var.df, it has just a wide shape. Both forms are useful for different computation and plotting requirements. For example, using the wide form wen can easily plot the relationship between the mean values in the red and blue channel with respect to the green one.
ggplot(vvm.obj$wide.var.df) +
geom_point(aes(x=green.avg.mean, y=red.mean/green.avg.mean), colour ='#BB7070') +
geom_point(aes(x=green.avg.mean, y=blue.mean/green.avg.mean), colour ='#7070BB') +
xlab('Green Avg mean [ADU]') +
ylab('Ratio of mean values to Green average [ADU]') +
labs(title='Ratio of mean values to Green average' ) +
theme(plot.title = element_text(size = rel(1.5)))
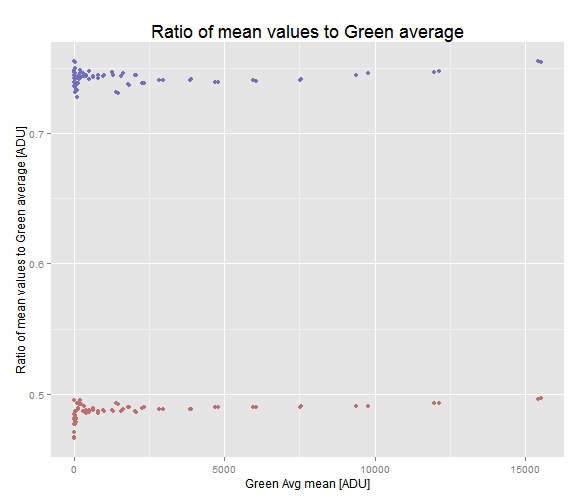
Ratio of red and blue mean values to the mean Green average. At higher exposition levels the ratios grow slightly.
Also, we may need to have the var.df and the cov.df information merged in a single data frame. We can get that through the merged.var.cov.df variable. However, this data frame can be expensive to build (requiring some seconds) and for that reason it is built at the variable first reading, subsequents use of these variable will just reuse the computed data without requiring a perceptible computing time.
head(vvm.obj$merged.var.cov.df, 8)
| pict | chan.a | chan.b | cov.a.b | mean.a | mean.b | var.a | var.b | channel.combo |
|---|---|---|---|---|---|---|---|---|
| _ODL5695.pgm | Red | Green R | 0.3391 | 696.1 | 1411 | 314.5 | 618.1 | Red, Green R |
| _ODL5695.pgm | Red | Green B | 36.1396 | 696.1 | 1415 | 314.5 | 597.3 | Red, Green B |
| _ODL5695.pgm | Red | Blue | 16.0167 | 696.1 | 1034 | 314.5 | 518.7 | Red, Blue |
| _ODL5695.pgm | Red | Green Avg | 18.2393 | 696.1 | 1413 | 314.5 | 593.5 | Red, Green Avg |
| _ODL5695.pgm | Green R | Green B | -14.2368 | 1410.8 | 1415 | 618.1 | 597.3 | Green R, Green B |
| _ODL5695.pgm | Green R | Blue | 4.7311 | 1410.8 | 1034 | 618.1 | 518.7 | Green R, Blue |
| _ODL5695.pgm | Green R | Green Avg | 301.9246 | 1410.8 | 1413 | 618.1 | 593.5 | Green R, Green Avg |
| _ODL5695.pgm | Green B | Blue | -19.0844 | 1414.6 | 1034 | 597.3 | 518.7 | Green B, Blue |
There are more components of the vvm class we haven't explained yet. But as they are very similar in both, the vvm and the hvdvm class, we will explain them in a later section common to both classes.
Digesting Samples After their Conversion to RGB
In the previous section we have seen how the vvm$digest() function process raw image samples and computes some statistics to analyze the raw image noise. Those statistics are in the same color space as the pixel values in the raw image, they are in the camera sensor raw color space. However, we don't want to finish the analysis just there, we want to track the noise to the final image. To allow this, there is a vvm$digest.as.rgb() function which converts the pixel values from the raw camera space to a chosen RGB space. For example, we can convert the pixel values to sRGB and then compute statistics over the pixels in that RGB space.
To make possible this conversion, from the camera raw to a RGB color space, we need some camera sensor color information. This camera color information can be easily obtained converting a camera raw image photo file to the .DNG (Adobe digital negative) format using the Adobe DNG Converter. The Windows version can be downloaded from here and the Mac OS X version from here. The Linux users will need the help of a friend converting any raw photograph. This color information very possible changes with the ISO speed, so you will need a conversion for each ISO you want to work with in imgnoiser.
The imgnoiser package contains as example the nikon.d7000.ISO100.colmap list, with the required information for a Nikon D7000 camera at ISO 100. You can use this list as a baseline and replace the data content according to your camera sensor.
str(nikon.d7000.ISO100.colmap)
#> List of 6
#> $ cam.matrices.1 :List of 5
#> ..$ illum.cct.k : num 2856
#> ..$ color.matrix : num [1:3, 1:3] 0.8642 -0.3999 -0.0453 -0.3058 1.1033 ...
#> ..$ forward.matrix : num [1:3, 1:3] 0.7132 0.2379 0.0316 0.1579 0.8888 ...
#> ..$ cam.calib.matrix: num [1:3, 1:3] 1 0 0 0 1 0 0 0 1
#> ..$ reduct.matrix : num [1:3, 1:3] 1 0 0 0 1 0 0 0 1
#> $ cam.matrices.2 :List of 5
#> ..$ illum.cct.k : num 6504
#> ..$ color.matrix : num [1:3, 1:3] 0.82 -0.487 -0.104 -0.224 1.239 ...
#> ..$ forward.matrix : num [1:3, 1:3] 0.6876 0.2676 0.0151 0.3081 1.0343 ...
#> ..$ cam.calib.matrix: num [1:3, 1:3] 1 0 0 0 1 0 0 0 1
#> ..$ reduct.matrix : num [1:3, 1:3] 1 0 0 0 1 0 0 0 1
#> $ analog.balance : num [1:3] 1 1 1
#> $ black.raw.level: num 0
#> $ white.raw.level: num 15892
#> $ tone.curve :'data.frame': 64 obs. of 2 variables:
#> ..$ src : num [1:64] 0 0.0159 0.0317 0.0476 0.0635 ...
#> ..$ dest: num [1:64] 0 0.0125 0.0411 0.0841 0.1326 ...
Once you get the .dng file, you can read the required information using ExifTool, that is a command line application which can be installed on Windows, Mac OS X and Unix systems. If you are note comfortable with this kind of application, there are also GUI interfaces taking advantage of the ExifTool engine, check the ExifTool web page.
For Windows I use ExifToolGUI, where the relevant data for the color transformation from raw to RGB is:
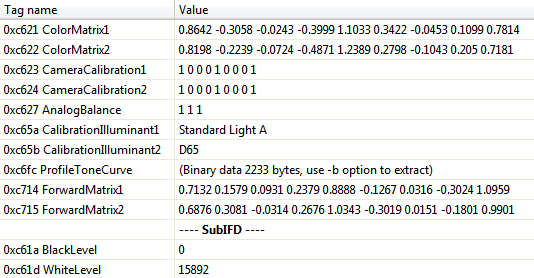
DNG Exif data from a Nikon-D700 at ISO 100. Also are required the reduction matrices (with tags 0xC625 and 0xC626).
It is also required the, reduction matrices with exif tags 0xC625 and 0xC626. In the case of the Nikon D7000 these tags don't exist in the .dng file, that's why they don't appear in the image above. So, I fill the corresponding entries in the list with 3x3 identity matrices. Once you get ready your camera color information in that R list, don't forget to save it for future use. If you kindly sent me the saved R object by email, I will gladly include it in the package.
The color information in that R list is needed for the creation of an object instance of the class colmap in imgnoiser:
cm.obj <- imgnoiser::colmap$new(imgnoiser::nikon.d7000.ISO100.colmap)
Now we need to let the colmap object to find out the white balance adjustment adequate for our image samples. This means that if our samples are not shots to a neutral surface, we will need to take a shot to a white object to make this white balance calibration.
To compute the white balance, we need the raw channel values of a camera raw reference white. This is like when in a photo processing application we white-balance the image by picking a neutral color area with an eyedropper. As the target in the image samples is a white surface, we can choose any of them, but if you look at the mean ratios in the image above, you will realize the color balance changes a little with the exposition level. Considering this, we will choose for our raw reference white, the images with Green Avg almost over 7,500 ADU, which is approximately at the middle point of their vertical variation range:
subset(vvm.obj$wide.var.df, abs(green.avg.mean - 7500) < 400)
| pict | red.mean | red.var | green.r.mean | green.r.var | green.b.mean | green.b.var | blue.mean | blue.var | green.avg.mean | green.avg.var | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 15 | _ODL5709.pgm | 3701 | 1864 | 7541 | 3492 | 7553 | 3827 | 5598 | 2843 | 7547 | 3785 |
| 16 | _ODL5710.pgm | 3691 | 1986 | 7523 | 3514 | 7539 | 3875 | 5577 | 3063 | 7531 | 3670 |
From this images we compute their mean color values:
sel <- subset(vvm.obj$wide.var.df, abs(green.avg.mean - 7500) < 400)
# Keep only the color mean columns
sel <- sel[,c('red.mean', 'green.avg.mean', 'blue.mean')]
# Get their means
apply(sel, 2, mean)
#> red.mean green.avg.mean blue.mean
#> 3696 7539 5588
We will use this color coordinates as the white reference argument in a call to the function get.conv.matrix.from.raw() in the cm.obj object we have just created. That raw white reference must be specified as a vector with the RGB color components.
The destination space, to.space, is the second parameter required in the call to the get.conv.matrix.from.raw() function. The possible values are:
sRGBAdobe RGBProPhoto RGBCIE RGB
We will use sRGB for our example:
neutral.raw <- c(3696, 7539, 5588)
raw.to.sRGB <- cm.obj$get.conv.matrix.from.raw(from.neutral.raw = neutral.raw, to.space = 'sRGB')
#> White Balance CCT:5360.34853090509
#> White Balance xy (1931):0.3359028328649360.348287791612151
# Lets print the conversion matrix
raw.to.sRGB
#> [,1] [,2] [,3]
#> [1,] 3.53864420 -0.6478555 -0.1170973
#> [2,] -0.35326743 1.6515491 -0.6454761
#> [3,] 0.03012944 -0.4927949 1.9944290
The get.conv.matrix.from.raw() function reports the white balance CCT found for the given white reference (5,360 °K in or example) and returns (in "invisible" way) the conversion matrix from camera raw to the target RGB space (sRGB in our example). If we want, we can receive and keep that result in an R variable to print it or do whatever else we want with it. But, as the package keeps this matrix, and other information, inside itself for the following color conversion, we don't have to do that.
Now we are almost ready to call the digest.as.rgb() class function, through our vvm.obj object, which makes the color conversion and computes the noise statistics in the resulting RGB images. The parameters for this function are the same we used before in the call to the vvm.obj$digest() function, in the previous section, plus additional ones regarding to the color conversion. One of them is the map.to.rgb parameter, to provide a colmap object with the color conversion know-how and already calibrated to white-balance the samples, it is the cm.obj in our example.
Furthermore, we have to choose the target scale with the rgb.scale parameter. This argument determines the range (from zero to this value) of the resulting RGB color values. We will choose 255 as a customary range for RGB color coordinates. However, the resulting color values will not be integers, they will be floating point numbers holding the complete precision of our calculations.
Moreover, with the rgb.labels parameter, we can choose the labels for the color values in the resulting data frame. By default they are (red , green , blue) but you can change them if you want, for example to your native language.
The parameter is.neutral is for a logical argument indicating if the samples come from shoots to a neutral target. If it is true, during the color conversion, the white-balance parameters previously established in the call to get.conv.matrix.from.raw, will be refined for each sample. This may be required to counterbalance the white balance variation we saw in the the mean ratios plot above: the white balance CCT is kept the same for all the samples, only the raw balance is adjusted. This white balance adjustment is only possible when the samples are neutral.
The very last step in the conversion process from raw to RGB, is the application of a tone curve. Actually, it is the application of two tone curves, one is the camera tone curve and the other is a gamma curve corresponding to the target RGB space. The camera tone curve specification is part of the data supplied in the colmap object creation. It can be a NULL value, meaning it is not required or wanted a camera tone curve. This camera tone curve is aimed to deal with the camera tone idiosyncrasies in order to get nicer images or with a particular style, this way you will see in the camera reviews, "this camera has a <choose a fancy adjective> contrast".
After the application of the camera tone curve, the gamma curve corresponding to the RGB target space is applied to the image. Every RGB target space has a corresponding gamma correction curve. For example the sRGB color space has a tone curve very close to 2.2. In this article we explain the basic idea behind a tone curve or gamma correction curve.
The digest.as.rgb() function has a couple of parameters to let you choose if you want to apply the camera tone curve and the RGB target space tone curve or not.
The use.camera.tc (use the camera tone curve) parameter is for a logical value indicating if the camera tone curve should be applied to the image samples or not. If you set this argument to TRUE —and you provided a tone curve in the colmap object creation data— it will be applied to the images, otherwise it wont be applied.
The target.tc parameter deals with the target RGB gamma correction. Its default value is yes, apply the target space tone curve. However, you can specify other value as:
linear: Do not apply a target space gamma curve at all.sRGB: Apply the sRGB tone curve.ProPhoto: Apply the ProPhoto tone curve.BT.709: Apply the ITU-R Recommendation BT.709 tone curve.Gamma.2.2: Apply a regular 2.2 gamma correction tone curve.Gamma.1.8: Apply a regular 1.8 gamma correction tone curve.- A data frame with two columns specifying the tone curve.
This allows to play and use other known color spaces. For example, it is said Adobe Lightroom internally uses the ProPhoto space with the sRGB tone curve. We can get that by setting ProPhoto as the target space in the call to the get.conv.matrix.from.raw() function and sRGB as the space tone curve in the call to digest.as.rgb().
As another example, we can set the digest.as.rgb() function parameters with use.camera.tc = FALSE and target.tc = 'linear' to get the pixel values in the RGB target space but keeping them linear.
The tone curve argument itself can be the tone curve. In such case, its first two columns must numeric and they must have at least 8 rows. The column names are not relevant. The first column will represent the source pixel value which will be mapped to the corresponding value in the second column. The values in the first column must contain the [0,1] range, and those in the second one must be in the [0,1] range. The missing values, the values between the given points of the curve, will be interpolated using smooth splines, with the required degree to pass over each data point.
As we are learning the use of the imgnoiser package and not really analyzing the noise data, in our current example, we will maintain the things simple don't using any tone curve and keeping the data linear.
To keep our previous noise statistics about the raw pixel values, we will create another vvm object, with the name rgb.vvm, to get the sRGB statistics, then we will plot the resulting sRGB linear statistics.
rgb.vvm <- vvm$new(has.RGGB.pattern = TRUE)
rgb.vvm$digest.as.rgb(file.name.from = '_ODL5695',
file.name.to = '_ODL5767',
file.name.ext = '.pgm',
file.path = img.path,
# Additional parameters for the color space conversion
is.neutral = TRUE,
map.to.rgb = cm.obj,
rgb.scale = 255,
target.tc = 'linear')
#> 70 image samples were successfully processed as RGBs.
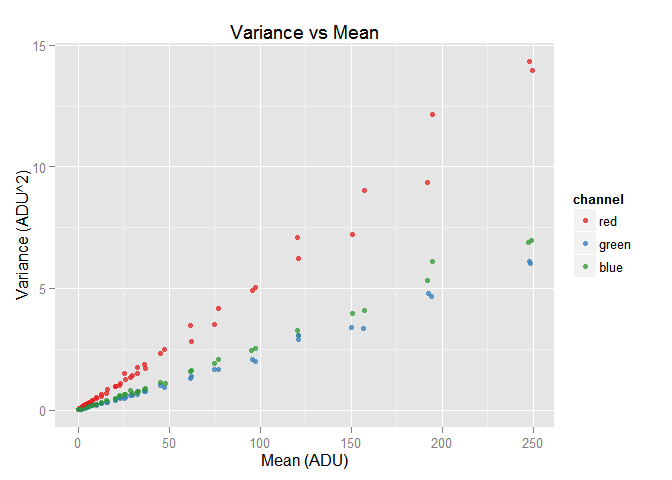
rgb.vvm$plot()
sorted.var.df <- dplyr::arrange(rgb.vvm$wide.var.df, green.mean)
sorted.var.df[54:57,]
| pict | red.mean | red.var | green.mean | green.var | blue.mean | blue.var | |
|---|---|---|---|---|---|---|---|
| 54 | _ODL5700.pgm | 37.24 | 1.710 | 37.27 | 0.7551 | 37.24 | 0.8848 |
| 55 | _ODL5701.pgm | 45.43 | 2.322 | 45.39 | 1.0159 | 45.44 | 1.1175 |
| 56 | _ODL5702.pgm | 47.73 | 2.473 | 47.67 | 0.9377 | 47.78 | 1.0942 |
| 57 | _ODL5703.pgm | 61.70 | 3.470 | 61.82 | 1.3105 | 61.72 | 1.5920 |
We can see the red component has the largest amount of noise among all the RGB the colors. A hint to explain this is the red raw values having a very lower scale than the others. From our raw white reference it is 3696/7539 = 49%: the half of the green scale. Also, the conversion matrix to build the red RGB color (first row) amplifies a lot the red raw component. However, as the focus here is to get used to the imgnoser package we won't get into more details.
Now we have similar data as we saw as result of calling the vvm.obj$digest() function, as the var.df, cov.df, wide.var.df and merged.var.cov.df data frames. Lets take a look at the last one.
head(rgb.vvm$merged.var.cov.df, 8)
| pict | chan.a | chan.b | cov | mean.a | mean.b | var.a | var.b | channel.combo |
|---|---|---|---|---|---|---|---|---|
| _ODL5695.pgm | red | green | -0.0949 | 22.65 | 22.62 | 1.0233 | 0.5003 | red, green |
| _ODL5695.pgm | red | blue | 0.0448 | 22.65 | 22.66 | 1.0233 | 0.5923 | red, blue |
| _ODL5695.pgm | green | blue | -0.1731 | 22.62 | 22.66 | 0.5003 | 0.5923 | green, blue |
| _ODL5696.pgm | red | green | -0.0995 | 23.36 | 23.38 | 1.1043 | 0.4748 | red, green |
| _ODL5696.pgm | red | blue | -0.0303 | 23.36 | 23.34 | 1.1043 | 0.5513 | red, blue |
| _ODL5696.pgm | green | blue | -0.3031 | 23.38 | 23.34 | 0.4748 | 0.5513 | green, blue |
| _ODL5697.pgm | red | green | -0.1031 | 28.92 | 28.93 | 1.3236 | 0.5961 | red, green |
| _ODL5697.pgm | red | blue | -0.0601 | 28.92 | 28.90 | 1.3236 | 0.8018 | red, blue |
As we have already said, there are more components of the vvm class we haven't explained yet. They are also available when we digest the images after their conversion to RGB with the digest.as.rgb() function. But as those components are very similar in both, the vvm and the hvdvm class we will explain them later in a section common to both classes.
Digesting Samples with the hvdvm Class
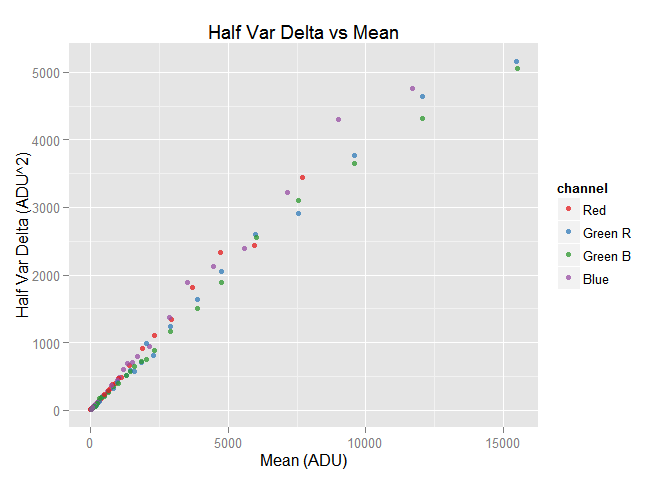
The hvdvm class computes the mean, and the half noise variance and covariance of the difference of two identical raw images. The theoretical basis of this approach is explained here and the practical application of that model is explained here.
Saving the Photographic Conditions in a .csv File
Compared to the vvm$digest() function, the hvdvm$digest() one, needs additional information to identify the shots taken under identical photographic conditions. To that effect, we have to prepare a .csv file relating our samples file names with the photographic conditions under which they were taken.
The .csv file must contain columns with the following names:
crop.file.namelightingISOshutter.speed(exposition time)aperturefocal.length
The letter case in the column names is not important. Also instead of period . you can use a white space. The columns sequence is not significant and there can be also other ones. Except for crop.file.name the exact values in these columns are not important, the hvdvm procedure will just distinguish the different values in them. In this sense, for example, shutter.speed can contain 1/50 or 0.02 or just 50. However, you must be consistent with your value coding and use the same value for the same condition and vice versa.
To build that .csv file, we will use the already mentioned ExifTool. This time using it —necessarily— as a command line application (not through a GUI interface): Open a command window on the folder containing the original raw image file photographs and run this command:
exiftool.exe -common -ext nef -csv ./ > photo-conds.csv
ExifTool will extract common (-common) exif metadata from the Nikon '.nef' image files (-ext nef) in the current directory (./) using the .csv format (-csv). The output redirection (> ISO100Conds.csv) will save the command output into the "photo-conds.csv" file in the current working directory. Of course, this works fine for Canon .CRW, .CR2 and many other proprietary file formats.
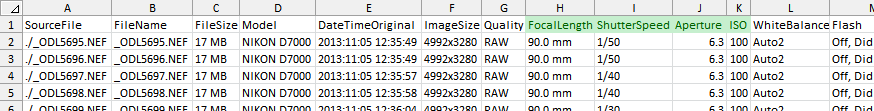
Now, we have to edit the .csv file produced by ExifTool to comply with the column naming and to add a column with the name of our sample files. If you open that file with MS Excel —annoyingly— it will convert the exposition times to dates (e.g 1/50 to Jan-50). To avoid that you can use LibreOffice. If you want to use MS Excel, rename the file to make it have a .txt extension, open it with MS Excel and convert the text lines to data with the "Text to Columns" menu command in the Data ribbon. Select the line with the column names and the data below it before to trigger the "Text to Columns" command. In the following dialog windows choose "delimited" by comma and assign the text type to the ShutterSpeed column. Also remove any other line in the file to just get the column titles in the first row and the photographic info below. After doing that I get what is shown below.
The columns required by imgnoiser have green background, you can remove the others if you want. Now rename, those columns to focal.length, shutter.speed and aperture.
If you want, you can leave the names as they come from ExifTool and set with the imgnoiser.option() function a map, using the 'conds.col.map' option, from your column names to the expected ones. The advantage of this approach is that you can reuse this mapping for countless .csv files and avoid renaming the column names every time. Also, the package options can be saved and reused in another R session. We will check that later.
We must add a column titled crop.file.name with the name of our samples using a formula taking as input the FileName column: take the first characters before the period . that separates the file name from its extension (NEF in my case) and concatenate the .pgm name extension in our sample files, something like =LEFT(C2,8)&".pgm". Copy that formula along all the rows.
Finally we need to add the lighting column with information based on our notes about the changes in the intensity of the light over the target scene. The exact values are not important. The class will be just interested on the different values in this column.
To begin with, you can fill all the column with the value 'Normal' and for those photos with higher lighting we can set the label 'High'. If you increased the lighting twice during the shooting session, you can correspondingly tag those photos with the values 'High A' and 'High B' in that column. Proceed with the same labeling schema for the shoots with lower lighting.
Use the "Save as..." command to save the file with the .csv format. Now we are ready to digest() the samples with the hvdvm class.
Digesting the Image Samples
We have to create an hvdvm object and call the digest() function giving as arguments the name of the .csv file we have just prepared and a path to the image samples referred from it:
hvdvm.obj <- hvdvm$new(has.RGGB.pattern = TRUE)
hvdvm.obj$digest('path/to/photo-conds.csv', 'path/to/images/')
You must notice the .csv file, in the crop.file.name column, contains only the sample file names, i.e. without the path to the folder containing them. That path is given in the second parameter call to the hvdvm$digest() function.
Now, we can plot the statistics obtained in the digestion:
hvdvm.obj$plot()
After the hvdvm digestion we have —again— those data frames we saw after using the vvm$digest() and vvm$digest.as.rgb() function: the var.df, cov.df, wide.var.df and merged.var.cov.df data frames contain the data extracted from the samples using the hvdvm technique. Lets take a look to wide.var.df:
head(hvdvm.obj$wide.var.df)
| cond | pict1 | pict2 | red.mean | red.var | green.r.mean | green.r.var | green.b.mean | green.b.var | blue.mean | blue.var |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | _ODL5709.pgm | _ODL5710.pgm | 3696 | 1814.2 | 7532 | 2910.6 | 7546 | 3103.7 | 5588 | 2394.8 |
| 2 | _ODL5707.pgm | _ODL5708.pgm | 2943 | 1347.1 | 6003 | 2592.8 | 6010 | 2560.2 | 4447 | 2127.0 |
| 3 | _ODL5705.pgm | _ODL5706.pgm | 2321 | 1111.9 | 4736 | 2047.9 | 4745 | 1884.3 | 3505 | 1883.0 |
| 4 | _ODL5703.pgm | _ODL5704.pgm | 1887 | 916.8 | 3862 | 1639.2 | 3869 | 1511.9 | 2864 | 1369.0 |
| 5 | _ODL5701.pgm | _ODL5702.pgm | 1417 | 662.8 | 2902 | 1234.3 | 2906 | 1172.0 | 2150 | 939.4 |
| 6 | _ODL5699.pgm | _ODL5700.pgm | 1124 | 484.8 | 2296 | 814.5 | 2300 | 881.8 | 1698 | 789.6 |
The pict1 and pict2 columns in the above data frame, and also present in other data frames in hvdvm objects, show the sample image file names to which the hvdvm procedure was applied: according with the data in the input .csv file, they come from photographs taken under identical conditions.
The first cond column, refers —with a sequential number— to the photographic conditions used to shot those samples. Different cond values represent different photographic conditions and vice versa.
We can get the different photographic conditions through the photo.conditions.df variable in the hvdvm objects.
# Get the different photographic conditions
head(my.hvdvm$photo.conditions.df)
| cond | lighting | iso | shutter.speed | aperture | focal.length | count |
|---|---|---|---|---|---|---|
| 1 | High | 100 | 1/10 | 6.3 | 90.0 mm | 2 |
| 2 | High | 100 | 1/13 | 6.3 | 90.0 mm | 2 |
| 3 | High | 100 | 1/15 | 6.3 | 90.0 mm | 2 |
| 4 | High | 100 | 1/20 | 6.3 | 90.0 mm | 2 |
| 5 | High | 100 | 1/25 | 6.3 | 90.0 mm | 2 |
| 6 | High | 100 | 1/30 | 6.3 | 90.0 mm | 2 |
Now we can see the photographic conditions referred by the column cond in the other hvdvm data frame variables. For example, the image samples in the first row in the wide.var.df variable, shown previously, come from photographs taken at an exposition time of 1/10s.
There are more components of the hvdvm class we haven't explained yet. But as they are very similar in both, the vvm and the hvdvm classes, we will explain them in a following section common to both of them.
Usage Common to the vvm and hvdvm Classes
In this section we will continue explaining the concepts and general workflow, common to the usage of the hvdvm or the vvm class. At this point, we will assume we have already digested some images with those classes.
The functions we will mention here can be called from a vvm or a hvdvm instance object. In the code excerpts we will use the one in the my.obj variable which can be an instance of any of those classes.
Introduction to Model Fitting
The notion of model in the imgnoiser package involves two concepts:
- Model Data Source: The specific data which will be used as the observations over which we will fit a model or regression.
- Model Family: The mathematical model or procedure allowing us to fit a model or regression over the data mentioned in the previous point.
When we fit a model in imgnoiser, we use a data source, fit a regression model available in a model family over that data, and give a name to that fitted model.
Model Data Source
Actually —by default— only one model data source exists and its named std.var. This model uses variables from the noise variance var.df result we get after calling the digest() function. In particular, in std.var data model the mean is the predictor variable (x) and var is the predicted one (y). This observations are grouped by channel.
You may add more data models at your will by changing the get.model.predictions option value.
When a function requires as an argument, the name of a model data source, by default, it will use the fit.model.data option value, which as you might guess, by default is std.var. Unless you provide more models, leave the default value for those arguments (typically named model.data.name ), as we will do in the following examples.
Model Family
We can choose, among different statistical regression models, the one that we want to use on our model data source. We group those models by the R function that actually makes the fitting.
By default, the following model families, from the stat package, are available:
lmlmrobsmooth.spline
As these fitting functions allow a wide variety of options, in fact directing them to different regression models or fitting procedures,
we have a broader amount of options than may it seem at first glance. As a trivial example, you can use the simple linear model lm() with weights.
When a function requires as an argument a model family, it will use by default the fit.model.data option value, which by default is 'lm'. You can add more fitting function at your taste by changing the fit.model.family option value.
Fit a Model
We can start using a simple (OLS: ordinary least squares) line fitting. To this effect we will just left the default value for all the fit.model() function arguments:
# Fit the default 'lm' model and save it with the default 'standard' name.
my.obj$fit.model()
#> The model "standard" was successfully fitted using:
#> lm(formula = var ~ mean + I(mean^2), data = model.src("std.var"))
Notice the output after the model fitting mentions the model family (lm), the formula (formula) used to relate predicted and predictor variables and the model source data (data = model.src("std.var")). If there would be any other parameter in the calling to the lm() function, they will appear in this output. Everything sent to the fitting functions will appear in this output; nothing is sent "under the hood".
Notice we did not input explicitly a formula in the call to the fit.model() function, it has been inferred from the source data model variables and the default argument for the degree parameter of that function, which for the hvdvm class is 1 and for the vvm one is 2. This argument indicates a linear relationship with the predicted variable var being predicted by a first or second degree polynomial on mean. If you set degree = 3, a cubic polynomial on mean will be fitted to predict the var variable, with 4 you will fit a bi-quadratic one and so on.
Instead of the degree you can directly set the formula argument for the fitting with the specific formula you want. In such cases, when you specify the formula, the degree argument is silently ignored. For example, we may want a regression predicting log2(var) using as predictor log2(mean), in such case we can fit the model using:
# Fit a model named 'log' using the 'lmrob' family:
my.obj$fit.model('log', formula = log2(var) ~ log2(mean), model.family = 'lmrob')
#> The model "log" was successfully fitted using:
#> lmrob(formula = log2(var) ~ log2(mean, data = model.src("std.var"))
The restrictions for the formula parameter are those from the model family function. In the previous example, the restrictions are from the `robustbase::lmrob()' function. For this reason, we can not use formulas with the 'smooth.spline' model family.
Other default arguments in the call to the fit.model() function we just did, is the name to assign to the model we are fitting (model.name parameter). Its default value is the fit.model.name option value, with standard as factory value.
Also, the confidence level (conf.level parameter) is used to compute the prediction interval limits. Its default value is the conf.level option value, with 95% as factory value.
We can call the fit.model() function again but with an argument for the model.name parameter different to standard to do not overwrite the model we just fitted, or we can omit again this argument to overwrite that model.
Lets fit a robust linear model using as weight the reciprocal squared values of the mean. We will assign the name 'weighted' to that model:
# This time we will set the model name and model family parameters
my.object$fit.model(model.name = 'weighted', model.family = 'lmrob', weights=1/mean^2)
#> The model "weighted" was successfully fitted using:
#> lmrob(formula = var ~ mean + I(mean^2), data = model.src("std.var"), weights = 1/mean^2)
Notice how we set the weights argument for the lmrob function in the call to fit.model() using the variable names in our model. This works because all the parameters that don't correspond to the fit.model() function are passed directly to the fitting function, lmrob in this case. This allows the use any and every parameter available for them.
Get a Fitted Model Predictions
We can get the predictions from the standard model we just fit, by calling the get.model.predictions() function with the default arguments:
# Get a data frame with the model predictions for the 'standard' model
preds <- my.obj$get.model.predictions()
head(preds,4)
tail(preds,4)
| mean | var | channel | fit.se | lcl | ucl |
|---|---|---|---|---|---|
| 5.317 | 2.937 | Red | 10.756 | -163.03 | 168.9 |
| 10.306 | 5.246 | Red | 10.716 | -160.71 | 171.2 |
| 19.976 | 9.723 | Red | 10.639 | -156.20 | 175.6 |
| 38.717 | 18.406 | Red | 10.494 | -147.47 | 184.3 |
| mean | var | channel | fit.se | lcl | ucl |
|---|---|---|---|---|---|
| 13079 | 6330 | Green Avg | 49.33 | 6013 | 6648 |
| 13896 | 6750 | Green Avg | 56.89 | 6426 | 7075 |
| 14713 | 7174 | Green Avg | 66.45 | 6839 | 7508 |
| 15529 | 7599 | Green Avg | 77.89 | 7251 | 7948 |
The columns in the resulting predictions data frame are:
mean: The predictor variable.var: The predicted variable.channel: The grouping factor.fit.se: The prediction standard error.lpl: The lower prediction interval limit.upl: The upper prediction interval limit.
The prediction interval is based on the confidence level argument (conf.level parameter), which by default is the same one used to fit the model. If you want technical details about these lasts columns, please check the R predict() function with respect to the level = conf.level, se.fit = TRUE and interval = 'prediction' arguments.
If we don't specify a model name, the predictions are computed using the default model, and the predictor variable is chosen for evenly spaced points in the same range as the source data used to fit the model. However, you can specify those predictor values and select the channels whose predictions you want.
For example, lets get the predictions from the 'weighted' model, for predictor values from 5000 to 8000 with steps of 1000, for the Blue and Red channels, with prediction intervals with 98% of confidence level:
# Get the predictions from the 'weighted' model and the given predictor values
my.obj$get.model.predictions( model.name = 'weighted',
x = seq(5000, 8000, 1000),
select = c('Blue','Red'),
conf.level = 0.98)
| mean | var | channel | fit.se | lpl | upl |
|---|---|---|---|---|---|
| 5000 | 2577 | Blue | 27.98 | 2125 | 3028 |
| 6000 | 3141 | Blue | 39.42 | 2597 | 3686 |
| 7000 | 3723 | Blue | 53.85 | 3084 | 4361 |
| 8000 | 4321 | Blue | 71.27 | 3586 | 5056 |
| 7000 | 3861 | Red | 103.6 | 3168 | 4555 |
| 8000 | 4532 | Red | 137.6 | 3722 | 5342 |
| 9000 | 5233 | Red | 176.7 | 4299 | 6166 |
| 0000 | 5963 | Red | 221.0 | 4898 | 7028 |
Get a Model Summary
To get a model summary, we call the print.model.summary() function indicating the name of the model whose summary we want. As usual, if we don't give a model name, the default model will be used:
# Get the default model summary
my.obj$print.model.summary()
#> #-------
#> # Model Name: "standard"
#> #
#> # Call:
#> # lm(formula = var ~ mean + I(mean^2), data = model.src("std.var"))
#> #-------
#>
#> ##-- channel : "Red" --##
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -273.30 -5.89 1.24 3.06 281.97
#>
#> Coefficients:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) 4.76e-01 1.08e+01 0.04 0.96
#> mean 4.63e-01 1.40e-02 32.95 < 2e-16
#> I(mean^2) 1.15e-05 2.15e-06 5.35 1.2e-06
#>
#> Residual standard error: 68.8 on 67 degrees of freedom
#> Multiple R-squared: 0.995, Adjusted R-squared: 0.995
#> F-statistic: 6.84e+03 on 2 and 67 DF, p-value: <2e-16
The print.model.summary() function —by default— prints the summary of each fitted channel in the model. You can use the select parameter to choose which specific models summaries you want. You can pick them by index number (between 1 and 4) or by channel label. For example, lets get the summary for the Blue channel in the weighted model:
my.obj$print.model.summary('weighted', select='Blue')
#> #-------
#> # Model Name: "weighted"
#> #
#> # Call:
#> # lmrob(formula = var ~ mean + I(mean^2), data = model.src("std.var"), weights = 1/mean^2)
#> #-------
#>
#> ##-- channel : "Blue" --##
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -335.108 -10.786 -0.235 5.168 467.468
#>
#> Coefficients:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) 2.40e+00 1.97e-01 12.19 < 2e-16
#> mean 4.73e-01 5.93e-03 79.79 < 2e-16
#> I(mean^2) 8.33e-06 1.45e-06 5.75 2.4e-07
#>
#> Robust residual standard error: 0.0375
#> Multiple R-squared: 0.996, Adjusted R-squared: 0.996
#> Convergence in 11 IRWLS iterations
This function calls the base::summary() function corresponding to the model family used for the model fitting. Those functions accepts other parameters you can also specify when calling to the print.model.summary() function.
For example, the summary.lm() function is used for models from the 'lm' model family, and this functions accepts the logical parameter correlation; we can pass this parameter through a call to the print.model.summary() function:
# The default model uses the 'lm' model family
my.obj$print.model.summary(select='Blue', correlation = TRUE)
Get the Fitted Model Object
The R functions that actually fit the models, like lm() or lmrob(), return an object which can be used in other R functions supporting them.
For example, the function car::avPlots() Added-Variable Plots, constructs and plots added-variable (also called partial-regression) models as an analysis tool to check the good fitness of each component in a linear model. We can use that function to find out if the linear and quadratic component mean component in the vvm models have a good fit. However, to be able to do that, we need the object returned by the fitting functions we mentioned above.
The function get.model() returns a list with the model objects for every fitted channels in a given model. As with the print.model.summary() function, we can select (select parameter) the channel(s) whose model we want, and as usual, if we don't provide a model name the default one will be used.
Lets get the Red model in the default model and use it to get the Added-Variable Plots:
red.fitted.object <- my.obj$get.model(select='Red')
car::avPlots(red.fitted.object\?span><span class="hl opt">)
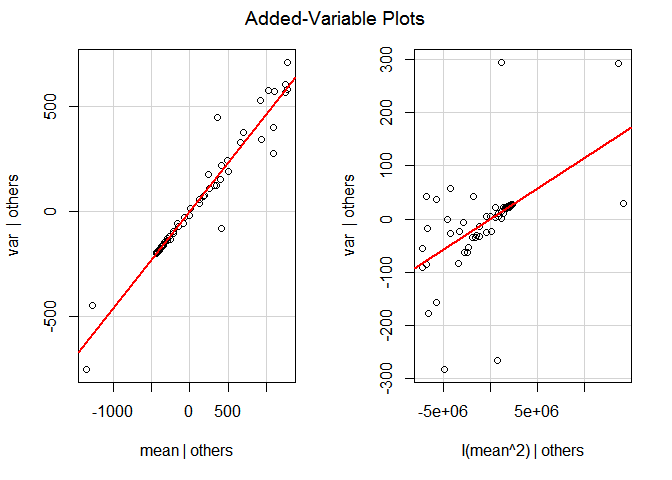
avPlots from the Red model object in the default ('lm') linear model. The quadratic member doesn't look too good.
Notice the get.model() function returns a list even when it is returning only one single object. That is the reason why we have to use [[1]] to extract the Red model object from the value returned by get.model() and pass it to the avPlots() function.
List, Query and Remove Fitted Models
The model.list variable provides a list with the already fitted models.
my.obj$model.list
#> $standard
#> [1] "lm(formula = var ~ mean + I(mean^2), data = model.src("std.var"))"
#>
#> $weighted
#> [1] "lmrob(formula = var ~ mean + I(mean^2), data = model.src("std.var"), weights = 1/mean^2)"
As you can see in the output above, the names in the list are the model names and the list components are the calls made to fit each model.
The exists.model() function tests if a model with a given name already exists. This function returns TRUE only if the model with the given name exists:
# We have a model named 'weighted'
my.obj$exists.model('weighted')
#> lmrob(formula = var ~ mean + I(mean^2), data = model.src("std.var"), weights = 1/mean^2)
#> [1] TRUE
# But we dont have other one called 'log-log'
my.obj$exists.model('log-log')
#> The model"log-log" does not exists.
#> [1] FALSE
We call the remove.model() function when we want to remove a fitted model. In this case, for security reasons, there is no default model name, you must specify it.
# We have a model named 'standard' ...
my.obj$exists.model('standard')
#> lm(formula = var ~ mean + I(mean^2), data = model.src("std.var"))
#> [1] TRUE
# .. but we don't want it any more
my.obj$remove.model('standard')
#> The model with name"standard"has been removed.
# Now, it doesn't exists
my.obj$exists.model('standard')
#> The model"standard" does not exists.
#> [1] FALSE
Plotting
There is nothing like a good plot to let the data speak by itself, showing us its features, validating our assumptions or raising new questions.
With the data we get through object variables like var.df, cov.df or photo.conditions.df and the result from the get.model.predictions() function, we can built any kind of plot we want. However, the function plot() provides an easy way to plot the data and a model already fitted.
By default, without any argument, the plot() function just plots the model observations. Using the model.name parameter, we can include a fitted model in the plot. As a convenience, if we set the model.name parameter to TRUE, the default model (the one with the name equal to thefit.model.name option value) is plotted.
# Plot the data
my.obj$plot()
# rebuild the previously deleted default model
my.obj$fit.model()
#> The model "standard" was successfully fitted using:
#> lm(formula = var ~ mean + I(mean^2), data = model.src("std.var"))
# Now, lets plot the default ('standard') model
my.obj$plot(model.name = TRUE)
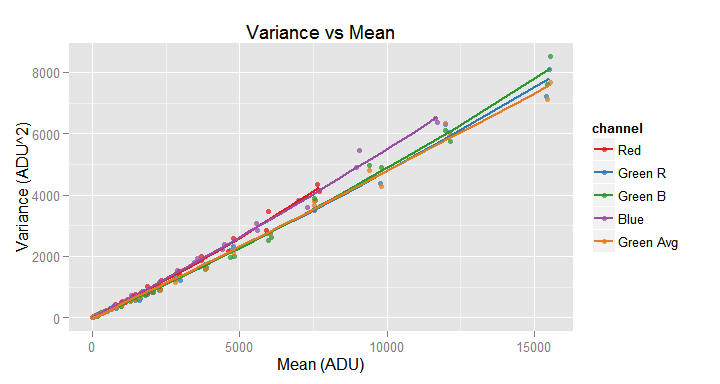
Adding the argument model.name = TRUE, the plot() function plots the data source observations and the default fitted model.
With the xlim and ylim parameters you can delimit the region to plot, zooming-in a particular area in the plot. Those parameters are vector of two numeric values with the desired x and y range for the plot.
Behind the scenes the ggplot2 package is drawing the plots. When we plot this kind of zoomed-in graphs, ggplot throws some warnings about the data being excluded from the plot because those axes ranges. The warnings logical parameter is FALSE by default to avoid them.
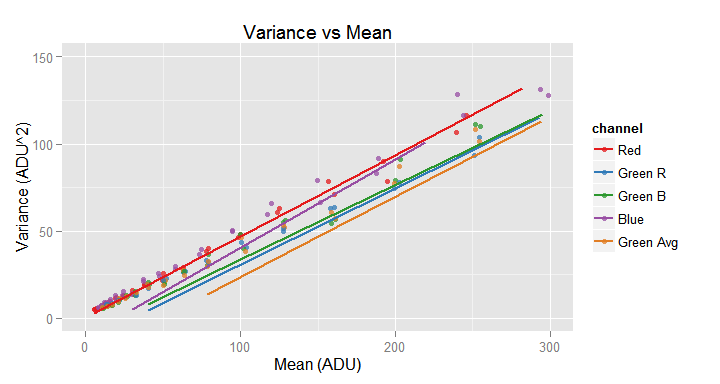
We may need to look closer some regions of a plot because the scale can hide interesting details. For example, if look closer to the axes origin, in the above plot, we get the following image.
# Get a closer look to the data points and the standard model in the axes origin
my.obj$plot(model.name = TRUE, xlim=c(0,300), ylim=c(0,150))
For the model.name parameter instead of TRUE (for the default plot), we can —of course— use the name of the fitted model we want to plot.
When we include a model in the plot, we can draw the fitted values (logical parameter fit) and the prediction intervals (logical parameter pred.int). By default, including a model in the plot causes to draw the fitted values as in the previous plots.
Lets plot the fitted values and the prediction intervals:
# Plot close-up to the data and the weighted model in the axes origin.
# Show the values fitted by the model and the prediction intervals.
# Mention in the sub title this is the weighted model
my.obj$plot(model.name = 'weighted', fit=TRUE, pred.int = TRUE,
xlim=c(0,300), ylim=c(0,150),
slab='Weighted Model')
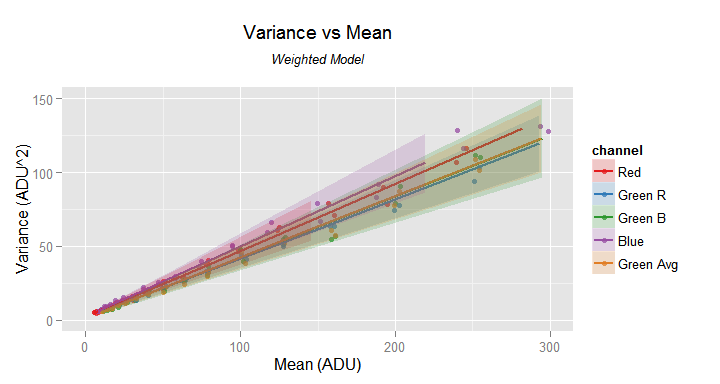
The 'weighted' model don't misses the data trend close to the axes origin. The prediction intervals are shown as shadowed areas. Notice the added sub-title.
In the previous image the prediction intervals are the semi-transparent areas around the lines through the fitted values. Be aware the prediction intervals are not available when the fitted model comes from the smooth.spline model family.
In the previous plot we have also added a subtitle: we can use the tlab parameter to set the main title and slab for the sub title. With the xlab and ylab parameters we can set the axes labels. If for any reason, you don't want any of those labels, set the corresponding parameter to NA. We will see a case for this in the following section.
The obs logical parameter determines if the data is shown (as points) in the plot or not, by default it is TRUE.
Additional plot customization ca be done with the following package options:
plot.line.width: The width of the lines passing through the fitted values.plot.point.size: The size of the points showing the source data.plot.point.opacity: The opacity of the points showing the source data.plot.ribbon.opacity: The opacity of the ribbons showing the prediction intervals.
The with parameter allows to specify a formula to filter the data that will be plot. For example, specifying with = ~ channel != 'Green Avg', will plot a graph excluding the data from the channel with the 'Green Avg' label.
We can use the x and y parameters to do some magic tricks. With then we can transform the fitted model and the source data to our wish. For example, lets say we want to plot the SNR (Signal to noise ratio) against the signal in a log-log (base 2) plot. In terms of our model variables, the SNR is the mean divided by the noise standard deviation, which is the squared root of the var variable (noise variance).
In other words, we want to plot the log2(mean) variable in the x axis and the log2(mean/sqrt(var)) in the y axis. Also, we should change the plot title because the default one, with these settings, wont correspond to the graph content anymore. In the following code example we will do this, also hiding the data points.
# Set the prediction interval ribbons to 10% opacity
imgnoiser.option('plot.ribbon.opacity', 0.1)
# Plot the SNR in a log-log graph
my.obj$plot(model.name = 'weighted',
x=log2(mean), y=log2(mean/sqrt(var)),
obs=FALSE, fit=TRUE, pred.int=TRUE,
tlab='Signal to Noise Ratio (SNR)',
slab='Weighted model')
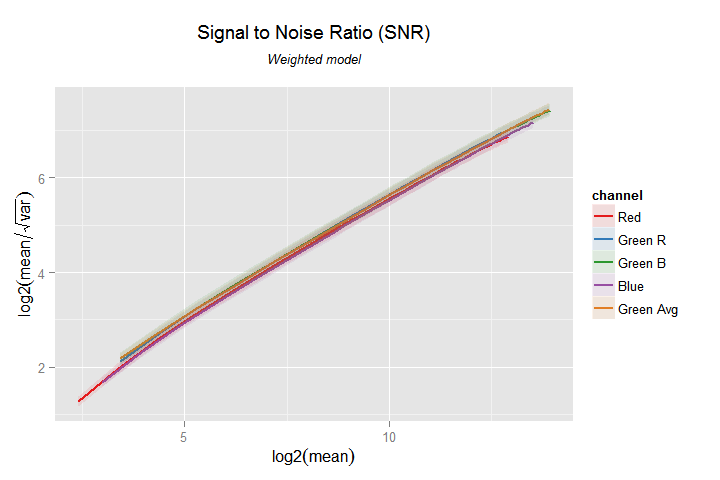
SNR by Signal level, in a log-log (base 2) graph. Notice the 'automatic' change of labels in the axes. We can customize them with the xlab and ylab parameters.
Notice how easily we have started the analysis processing the image samples and finished within minutes with a nice plot, showing even the prediction intervals.
Using the ggplot Object Returned by the Plot Function
Despite the flexibility of the plot() function, it is not unlikely you will need to cook the plot more with your own hands. To this effect, the plot() function returns the ggplot object it constructs to render the plot. However, this is an invisible return. You must receive it in a variable and use it the way you want.
The print parameter in the plot() function supports this idea by allowing us to set it to FALSE instructing the plot() function to build and pass the ggplot object but without showing or printing the plot. This way we can get that ggplot object, customize it more to our needing and finally print it.
For example, lets say we want to show side by side the plots of the 'standard' and the 'weighted' models close to the axes origin, sharing the same title, legend and y axis using this code which requires the ggplot object of those model plots:
# Get the standard linear regression plot
std.plot <- my.objj$plot(model.name=TRUE, print=FALSE,
xlab='Mean', ylab=NA, tlab=NA, xlim=c(0,300), ylim=c(0, 150))
# Get the weighted, robust, linear regression plot
weighted.plot <- my.obj$plot(model.name='weighted', print=FALSE,
xlab='Mean', ylab=NA, tlab=NA, xlim=c(0,300), ylim=c(0, 150))
# Use both plot objects for detailed customization.
shared.legend(std.plot,
weighted.plot,
'The Standard and Weighted Models',
'Variance (ADU^2)'
)
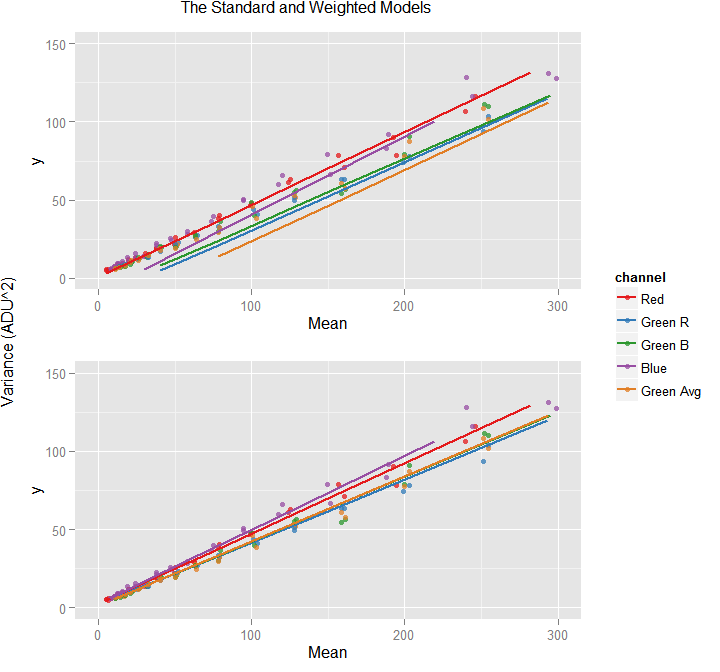
This is a single graph showing two plots built from the ggplot objects returned by the plot() function. This function constructed, one by one, the top and bottom plot, with settings avoiding to have title and y label.
In the example above, there is an implicit shared.legend() function which based on the ggplot of two graphs, puts them one above the other sharing a common title and y axis label. Correspondingly, the 'standard' and 'weighted' plots were built, but not printed, avoiding to have title and y label.
Persistence
With the imgnoiser package you can save and load:
- Object Instances
- The whole set of package options.
Saving and Loading a Class Object
With the imgnoiser.save() function we can save an object instance of the hvdvm or the vvm class to a disk file, and later —maybe in another R session or computer— retrieve it with the imgnoiser.load() function. This way you can share your data and findings with a friend or a community. The files are saved to disk have .imn as name extension.
If when saving an object, there is a file with the same name, it wont be overwritten unless the argument stop.overwrite = FALSE is specified.
By default, when we save an object, we also save in the same file all the imgnoiser package options. This makes sense because through the package options you can not only customize cosmetic or user interface preferences, but procedures in the form of functions, like fitting model procedures, source data model extraction, fitted model prediction functions and so on. You can change those options in such a way that the saved data (and meta-data) may be not enough to reproduce the environment when you save your noise analysis objects with the package.
When we load an object, by default, we also load all the imgnoiser package options saved with the object. The save.options and load.options logical parameters in the imgnoiser.save() and imgnoiser.load() functions allows to change that behavior and save or load the package options only when desired. Besides, you can save your current package options (we will see this in the following section) before to load the options with an object.
As a rule of thumb it is recommended to save the package options with the objects, to be able to load those options, or not, when you load the object.
As a convenience, if when saving an object we don't provide a file name, the object name will be used. This way, for example, the my.vvm.object will be saved in the 'my.vvm.object.imn' file. Also, if the file name does not have the .imn extension that will be automatically appended to it.
# List the models in the object
my.obj$model.list
#> $weighted
#> [1] "lmrob(formula = var ~ mean + I(mean^2), data = model.src(\"std.var\"), weights = 1/mean^2)"
#>
#> $standard
#> [1] "lm(formula = var ~ mean + I(mean^2), data = model.src(\"std.var\"))"
# Save the my.obj object, and the package options, in a file named 'my.obj.imn'
imgnoiser.save(my.obj)
#> The "my.obj" object was successfully saved in the "my.obj.imn" file.
# Destroy this objects to "make sure" we load the disk instance
my.obj <- NULL
from.disk.obj <- NULL
# Some days later, from another computer, load from disk the saved object
from.disk.obj <- imgnoiser.load('my.obj')
#> An instance of the "vvm" class was successfully load from the "my.obj.imn" file.
# List the models in the object
from.disk.obj$model.list
#> $weighted
#> [1] "lmrob(formula = var ~ mean + I(mean^2), data = model.src(\"std.var\"), weights = 1/mean^2)"
#>
#> $standard
#> [1] "lm(formula = var ~ mean + I(mean^2), data = model.src(\"std.var\"))"
Other saving and loading examples:
# Save the vvm.obj object in a particular path and file name with
# the package options. If there is afile with the same name just overwrite it
imgnoiser.save(vvm.obj, 'path/to/d7000-iso100-neutral', stop.overwrite = FALSE)
# Load the object but not the package options
hvdvm.obj <- imgnoiser.load('path/to/my.object', load.options = FALSE)
Saving and Loading the package Options
It is annoying to discover and set carefully the package options that suits you best, just to start all over again in the next R session.
With the imgnoiser.options.save() and imgnoiser.options.load() functions you can save and later load the whole set of package options.
# Save the options
imgnoiser.options.save('2015-04-12-options', stop.overwrite = FALSE)
# Load the options
imgnoiser.options.load('2015-04-12-options')
Conclusions
The imgnoiser package is a set of tools, concepts and definitions that helps to analyze image noise and makes easier to share the findings without delivering and explaining once and again the software environment, definitions and instructions to reproduce most of the results.